The NPFFR1 (neuropeptide FF receptor 1) responds to endogenous RF-amide peptides like NPFF (Neuropeptide FF) and RFRP-3 (RF-amide related peptide-3), both containing a conserved C-terminal Arg-Phe-NH2 motif (Quillet et al., 2016). In mammals, NPFF-related peptides are derived from two precursors (Bonini et al., 2000): pro-NPFFA, yielding peptides such as NPFF and NPAF (Neuropeptide AF) that prefer binding to NPFFR2 (neuropeptide FF receptor 2); and pro-NPFFB, yielding RFRP-3, which favors NPFFR1 (Quillet et al., 2016). NPFFR1 primarily couples to inhibitory Gi/o proteins and regulates diverse physiological functions, including energy homeostasis, reproduction, substance abuse disorders, cardiovascular control, anxiety, food intake, and pain (Nguyen et al., 2020).
Previous studies have suggested that NPFF receptors and their endogenous ligands are involved in the regulation of pain perception and opioid-induced antinociception (Ayachi and Simonin, 2014). The two receptors (NPFFR1 and NPFFR2) may exert opposing pain modulation effects, which may be attributed to the functional differences between NPFFR1 and NPFFR2. While NPFFR1 may act as an anti-opioid modulator, NPFFR2 may instead exert antinociceptive and opioid-potentiating effects (Malin et al., 2015). Blocking NPFFR1 alone can prevent analgesic tolerance and dependence, enhance opioid antinociceptive effects, and reduce opioid withdrawal symptoms. However, due to the lack of highly selective agonists and antagonists of NPFFR1 vs NPFFR2, their precise mechanism of opioid modulation is yet to be unraveled.
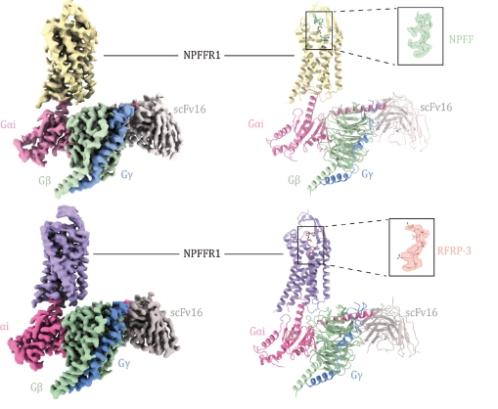
RFRP-3 and NPFF are endogenous ligands of NPFFR1, both sharing the four-amino acid sequence PQRF at their C-termini (Fig. 1A and 1B). However, the remaining four amino acids at the N-termini of each peptide result in different potencies at NPFFR1. This was confirmed through GloSensor assay (to measure the cAMP levels) using 293 T cells expressing NPFFR1 to assess the potency of these two ligands (Fig. 1C). While both ligands activate the receptor’s Gi signaling pathway, RFRP-3 exhibit ∼20-fold increase of potency to NPFFR1 compared to NPFF (EC50 = 1.99 nmol/L and 39.8 nmol/L for RFRP-3 and NPFF, respectively; Table S1). To investigate the underlying mechanisms of ligand recognition (the structural differences for the two RF-amide peptides bound to NPFFR1) and subtype-selectivity (between NPFFR1 and NPFFR2), we carried out structural studies for the two ligand-bound NPFFR1-Gi complexes. We designed the NPFFR1 construct as described in the Methods section. Cryo-EM maps were acquired at 3.16 Å and 3 Å global resolution for the NPFF-NPFFR1-Gi complex and RFRP-3-NPFFR1-Gi complex, respectively (Figs. 1D, 1E, S2 and S3; Table S2), and the atomic models were built and refined according to the cryo-EM maps. Local refinement of the receptor region yielded maps at 4.02 Å and 3.77 Å resolution for the NPFF-bound and RFRP-3-bound NPFFR1 structures, respectively.
Link to Publisher’s page: Molecular recognition at the opioid-modulating neuropeptide FF receptor 1 | Protein & Cell | Oxford Academic